




Optical Imaging Techniques and Technologies
I am a firm believer in the synergy between technology development and scientific discovery. The desire to answer interesting scientific questions is often the best driving force behind the creation of new technologies. At the same time, the development of novel scientific tools and techniques often enables us to ask new scientific questions. To that end, my research has focused on both the development of new techniques and technologies, as well as their application; specifically to investigate the details of neurovascular architecture.
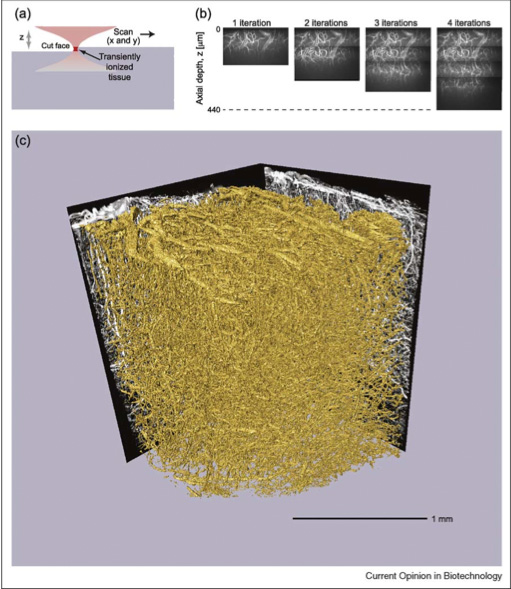
The main focus of my Ph.D. thesis was the invention of a technique that we call All-Optical Histology. The technique combines two-photon laser scanning microscopy and plasma-mediated femtosecond pulse laser ablation to perform serial iterations of imaging and ablation in histological tissue samples. In contrast to traditional reconstruction by serial thin sections, all-optical histology images are taken within an undisturbed block of tissue, and surface removal of the previously imaged tissue is performed by non-thermal laser ablation, which produces no sheer forces upon the soft tissue block. Thus, we obtain a fully-registered 3-dimensional reconstruction of a macroscopic block of tissue with micrometer resolution, without the tissue distortion and mis-registration issues inherent to serial section reconstruction.
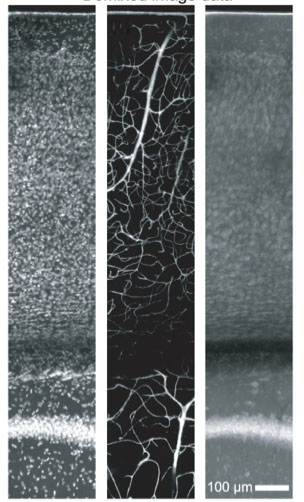
Tissue Staining and Clearing
In the course of applying our All-Optical Histology technique to the study of angio-architecture, we developed a variety of staining and clearing protocols in order to facilitate 3D reconstruction of of neurovascular networks. These techniques included the clearing of brain tissue with a graded mixture of Triton X-100 and sucrose to both index-match and minimize tissue distortion from osmotic shrinkage. We also developed perfusion techniques involving a mixture of FITC-albumin and gelatin, which allowed complete and stable fluorescent vascular fills that were formaldehyde-fixation stabilized and compatible with optical imaging of the surrounding neuronal tissue. We also demonstrated that, counter to conventional wisdom, deep staining of thick tissue (1-millimeter or more thickness) with directly-conjugated antibodies did indeed provide sufficient labeling contrast under two-photon microscopy.
The development of novel modes of microscopy necessitates the development of software tools to run the custom-built microscopes, collect data, and analyze the resulting image stacks to produce quantitative results. We developed MPScope, and later MPScope2*, to incorporate script-based automation for the collection of multiple-field histological data. We are currently developing MPScope3**, which will include accommodation of large field-of-view data acquisition, and standardized HDF5 data storage format.
The quantitative analysis of large 3D volumes of vascular and neuronal image data necessitated the development of a customized software package for the segmentation and vectorization of 3D data with marginal signal-to-noise. We developed the VIDA*** (Volumetric Image Data Analysis) suite of software to produce a radii-accurate vectorization of vascular networks and automated 3D localization of neuronal cell nuclei, including the use of a "Threshold Relaxation" algorithm to bridge occasional gaps in the vascular segmentation due to local variations in the signal-to-noise ratio of the underlying data.
*MPScope2 Primary developer : Quoc Nguyen, **MPScope3 Primary developer : Jonathan Driscoll, ***VIDA suite co-developer : John Kaufhold.
 Optical imaging deep
within tissue presents several technical challenges.
Local variations in the index of refraction within the
tissue (e.g., from membrane to protein, to intracellular
cytosol), results in the progressive scattering of
light. Meanwhile, the difference between the mean
refractive index of the tissue (reports vary from n = 1.35
to n = 1.4) and the refractive index of the immersion fluid
(typically water at n = 1.33, or oil at n = 1.52),
results in increasing spherical aberration with depth.
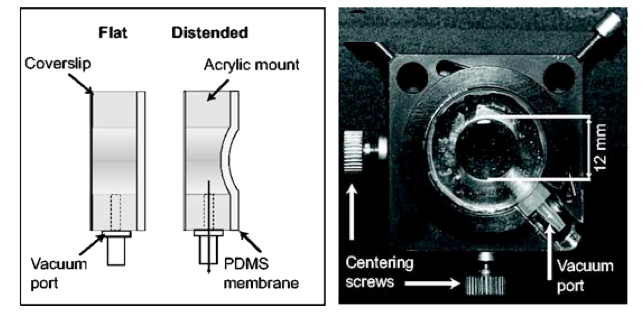
To counteract this second effect, we developed a novel
deformable membrane lens which utilizes air pressure to
produce a controllable membrane curvature. When placed
directly above the objective, the deformable lens allows the
continuous correction of spherical aberration with
increasing imaging depth into tissue.
Optical imaging deep
within tissue presents several technical challenges.
Local variations in the index of refraction within the
tissue (e.g., from membrane to protein, to intracellular
cytosol), results in the progressive scattering of
light. Meanwhile, the difference between the mean
refractive index of the tissue (reports vary from n = 1.35
to n = 1.4) and the refractive index of the immersion fluid
(typically water at n = 1.33, or oil at n = 1.52),
results in increasing spherical aberration with depth.
To counteract this second effect, we developed a novel
deformable membrane lens which utilizes air pressure to
produce a controllable membrane curvature. When placed
directly above the objective, the deformable lens allows the
continuous correction of spherical aberration with
increasing imaging depth into tissue.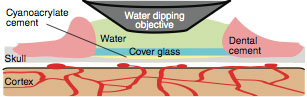 Optical access to neuronal tissue and neurovasculature is
typically obtained in one of two ways: 1) A complete
removal of bone from a millimeter-scale region of the skull,
i.e., a craniotomy, or 2) A thinning of the skull to
tens-of-micrometers thickness over a region ~100 micrometers
in size. While the thinned-skull preparation avoids
direct exposure of the brain, it is typically limited to
small imaging windows, and requires occasional re-thinning
of the skull to counteract bone regrowth. The
craniotomy provides a larger field of view, often with
agarose and coverslip reinforcement; but full exposure of
the brain opens the potential for inflammation and secondary
effects including spreading depression and vascular
re-organization.
Optical access to neuronal tissue and neurovasculature is
typically obtained in one of two ways: 1) A complete
removal of bone from a millimeter-scale region of the skull,
i.e., a craniotomy, or 2) A thinning of the skull to
tens-of-micrometers thickness over a region ~100 micrometers
in size. While the thinned-skull preparation avoids
direct exposure of the brain, it is typically limited to
small imaging windows, and requires occasional re-thinning
of the skull to counteract bone regrowth. The
craniotomy provides a larger field of view, often with
agarose and coverslip reinforcement; but full exposure of
the brain opens the potential for inflammation and secondary
effects including spreading depression and vascular
re-organization. We developed the PoRTS* (Polished, Reinforced Thinned Skull) Window surgical technique to garner the best of both worlds. A large region of the skull (several millimeters across) is thinned to tens-of-micrometers in thickness. We then fit and glue a coverslip segment into the indentation to provide mechanical support over the large thinned region and prevent regrowth of the bone. The result is a large-field optical window that requires no surgical maintenance, circumvents inflammation artifacts, and provides stable in vivo imaging from Day 0 out to Day 60 and beyond.
*PoRTS concept co-inventors : Patrick Drew, Andy Shih, David Kleinfeld.
manuscript authors: P. J. Drew, A. Y. Shih, P. Knutsen, D. Davalos, P. Blinder, K. Akassoglou, P. S. Tsai and D. Kleinfeld